Variance homogeneity in two or more groups
TODO
- link to variance
Install required packages
wants <- c("car", "coin")
has <- wants %in% rownames(installed.packages())
if(any(!has)) install.packages(wants[!has])Compare two groups
Boxplot with added stripchart
set.seed(123)
P <- 2
Nj <- c(50, 40)
DV1 <- rnorm(Nj[1], mean=100, sd=15)
DV2 <- rnorm(Nj[2], mean=100, sd=13)
varDf <- data.frame(DV=c(DV1, DV2),
IV=factor(rep(1:P, Nj)))boxplot(DV ~ IV, data=varDf)
stripchart(DV ~ IV, data=varDf, pch=16, vert=TRUE, add=TRUE)
\(F\)-test for variance ratio in two groups
var.test(DV1, DV2)var.test(DV ~ IV, data=varDf)
F test to compare two variances
data: DV by IV
F = 1.5437, num df = 49, denom df = 39, p-value = 0.1632
alternative hypothesis: true ratio of variances is not equal to 1
95 percent confidence interval:
0.8361247 2.7913772
sample estimates:
ratio of variances
1.543726 Mood-test for two groups (nonparametric)
mood.test(DV ~ IV, alternative="greater", data=varDf)
Mood two-sample test of scale
data: DV by IV
Z = 1.8065, p-value = 0.03542
alternative hypothesis: greaterAnsari-Bradley-test for two groups (nonparametric)
ansari.test(DV ~ IV, alternative="greater", exact=FALSE, data=varDf)
Ansari-Bradley test
data: DV by IV
AB = 1025, p-value = 0.02116
alternative hypothesis: true ratio of scales is greater than 1library(coin)
ansari_test(DV ~ IV, alternative="greater", distribution="exact", data=varDf)
Exact Ansari-Bradley Test
data: DV by IV (1, 2)
Z = -2.0304, p-value = 0.02138
alternative hypothesis: true mu is less than 1Compare more than two groups
Boxplot with added stripchart
Nj <- c(22, 18, 20)
N <- sum(Nj)
P <- length(Nj)
levDf <- data.frame(DV=sample(0:100, N, replace=TRUE),
IV=factor(rep(1:P, Nj)))boxplot(DV ~ IV, data=levDf)
stripchart(DV ~ IV, data=levDf, pch=20, vert=TRUE, add=TRUE)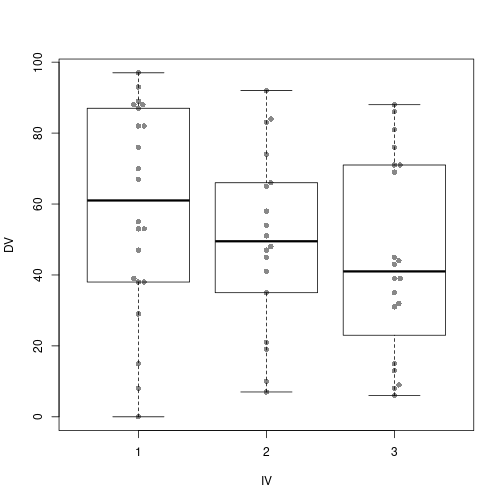
Levene-test
library(car)
leveneTest(DV ~ IV, center=median, data=levDf)Levene's Test for Homogeneity of Variance (center = median)
Df F value Pr(>F)
group 2 0.1456 0.8648
57 leveneTest(DV ~ IV, center=mean, data=levDf)Levene's Test for Homogeneity of Variance (center = mean)
Df F value Pr(>F)
group 2 0.0961 0.9085
57 Fligner-Killeen-test
fligner.test(DV ~ IV, data=levDf)
Fligner-Killeen test of homogeneity of variances
data: DV by IV
Fligner-Killeen:med chi-squared = 0.0936, df = 2, p-value = 0.9543library(coin)
fligner_test(DV ~ IV, distribution=approximate(B=9999), data=levDf)
Approximative Fligner-Killeen Test
data: DV by IV (1, 2, 3)
chi-squared = 0.0936, p-value = 0.9586Detach (automatically) loaded packages (if possible)
try(detach(package:car))
try(detach(package:coin))
try(detach(package:survival))
try(detach(package:splines))Get the article source from GitHub
R markdown - markdown - R code - all posts